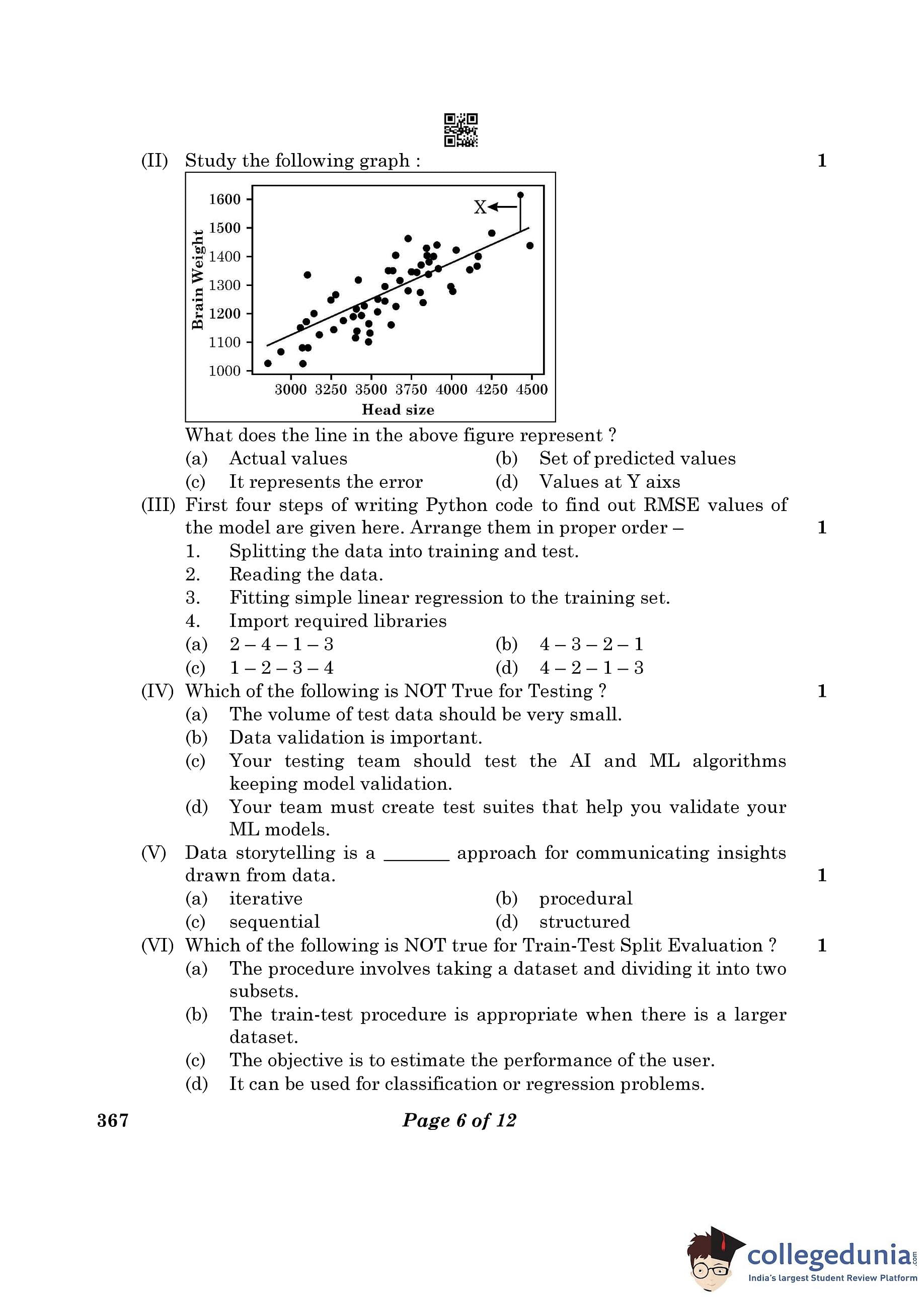
Identifying common transcriptome signatures of cancer by interpreting deep learning models, Genome Biology
Por um escritor misterioso
Last updated 16 junho 2024
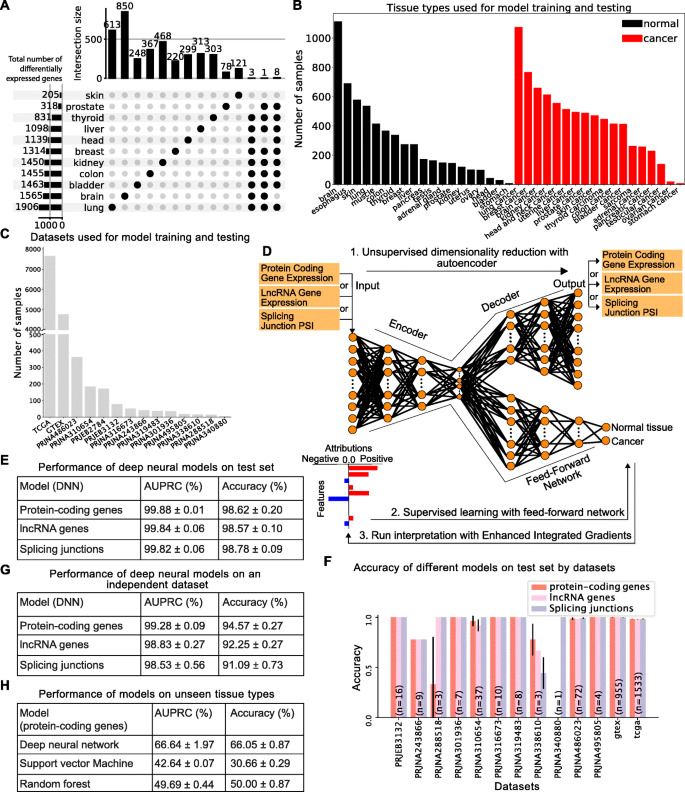
Background Cancer is a set of diseases characterized by unchecked cell proliferation and invasion of surrounding tissues. The many genes that have been genetically associated with cancer or shown to directly contribute to oncogenesis vary widely between tumor types, but common gene signatures that relate to core cancer pathways have also been identified. It is not clear, however, whether there exist additional sets of genes or transcriptomic features that are less well known in cancer biology but that are also commonly deregulated across several cancer types. Results Here, we agnostically identify transcriptomic features that are commonly shared between cancer types using 13,461 RNA-seq samples from 19 normal tissue types and 18 solid tumor types to train three feed-forward neural networks, based either on protein-coding gene expression, lncRNA expression, or splice junction use, to distinguish between normal and tumor samples. All three models recognize transcriptome signatures that are consistent across tumors. Analysis of attribution values extracted from our models reveals that genes that are commonly altered in cancer by expression or splicing variations are under strong evolutionary and selective constraints. Importantly, we find that genes composing our cancer transcriptome signatures are not frequently affected by mutations or genomic alterations and that their functions differ widely from the genes genetically associated with cancer. Conclusions Our results highlighted that deregulation of RNA-processing genes and aberrant splicing are pervasive features on which core cancer pathways might converge across a large array of solid tumor types.

Splicing signature database development to delineate cancer

Transcriptomics based multi-dimensional characterization and drug
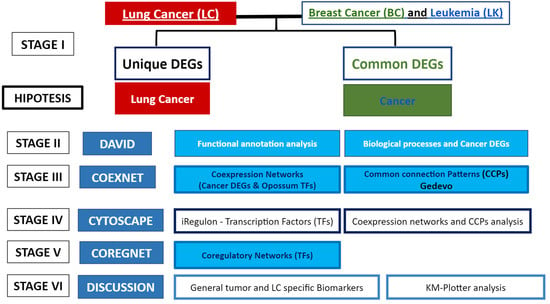
Biology, Free Full-Text
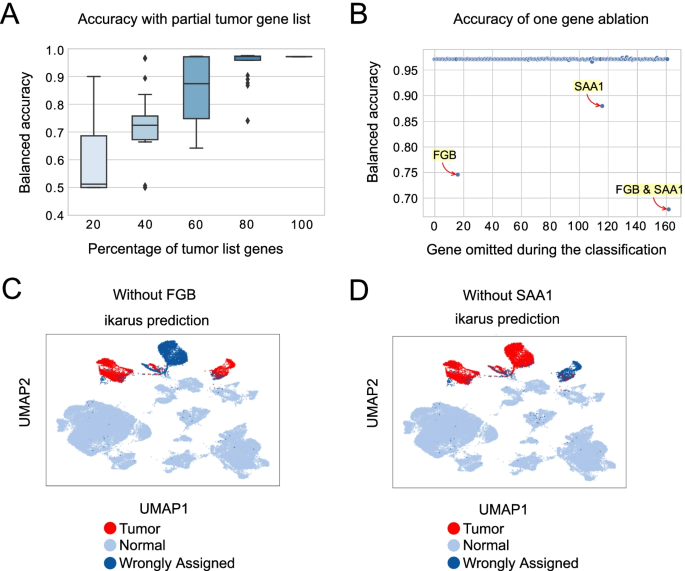
Identifying tumor cells at the single-cell level using machine
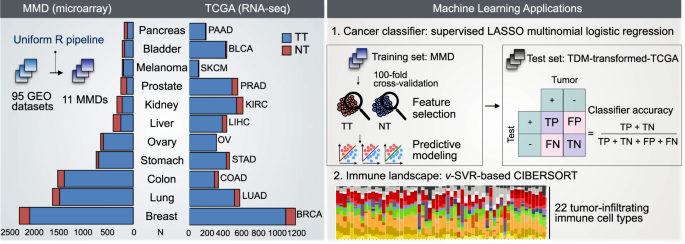
Compendiums of cancer transcriptomes for machine learning

A Comparative Analysis of Single-Cell Transcriptome Identifies
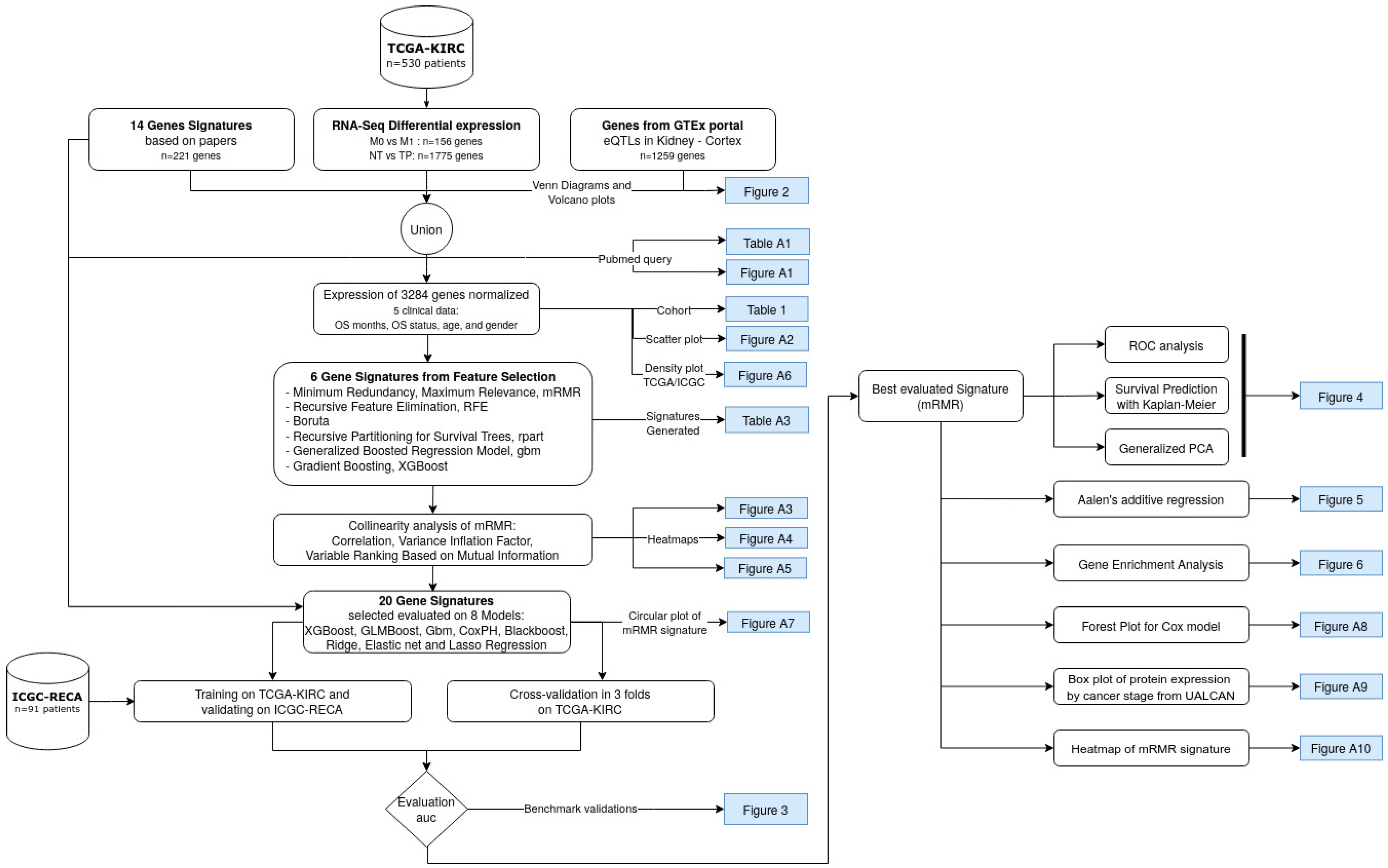
Cancers, Free Full-Text
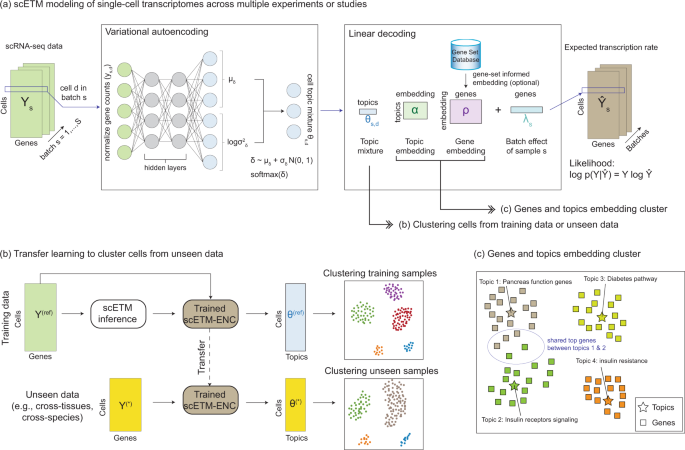
Learning interpretable cellular and gene signature embeddings from
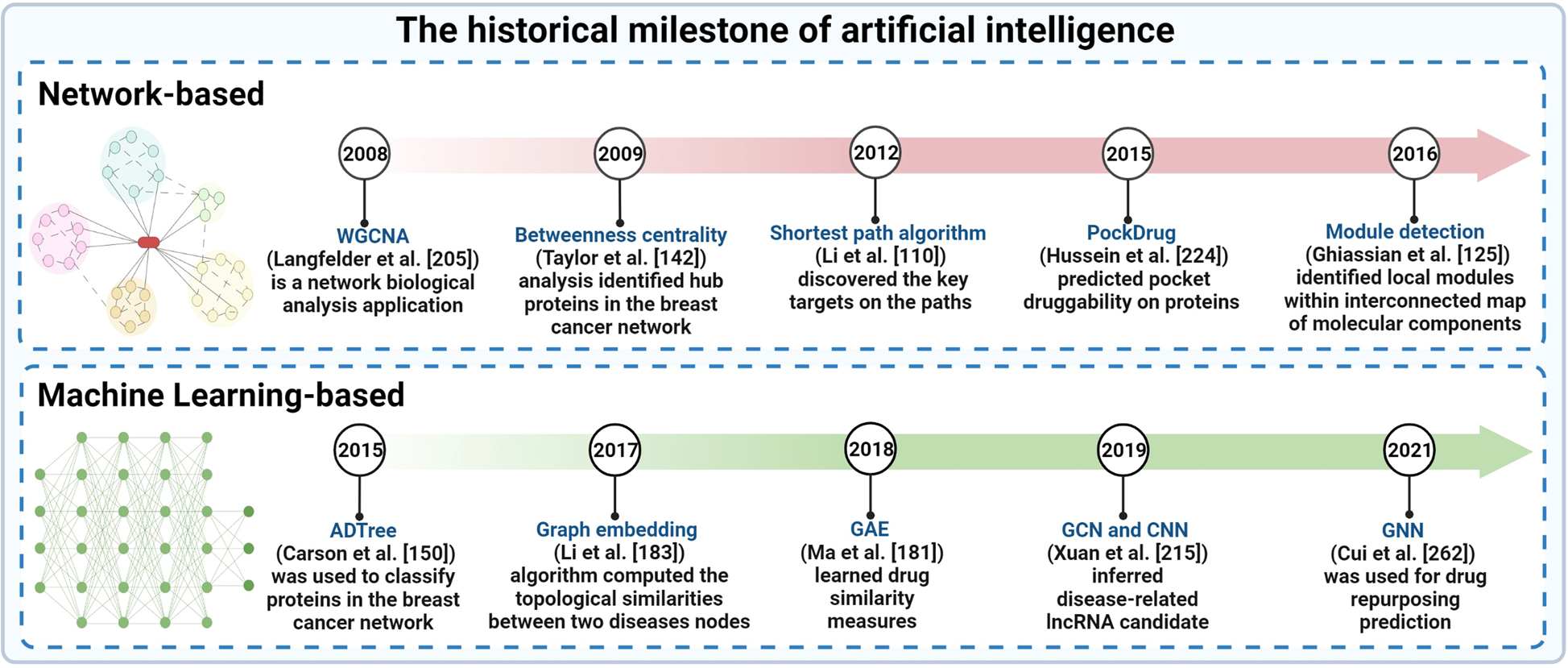
Artificial intelligence in cancer target identification and drug
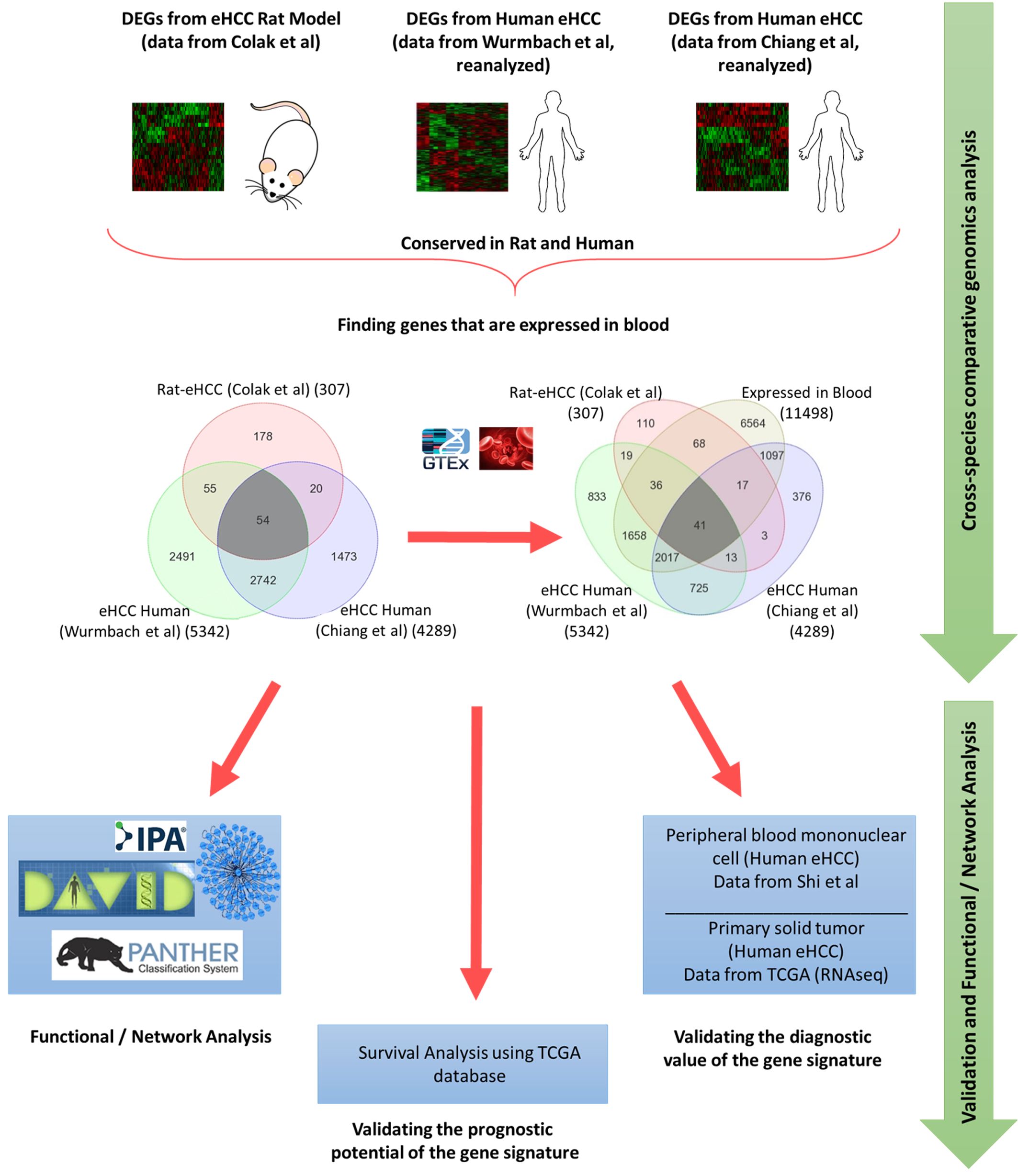
Frontiers Identification of Gene Signature as Diagnostic and
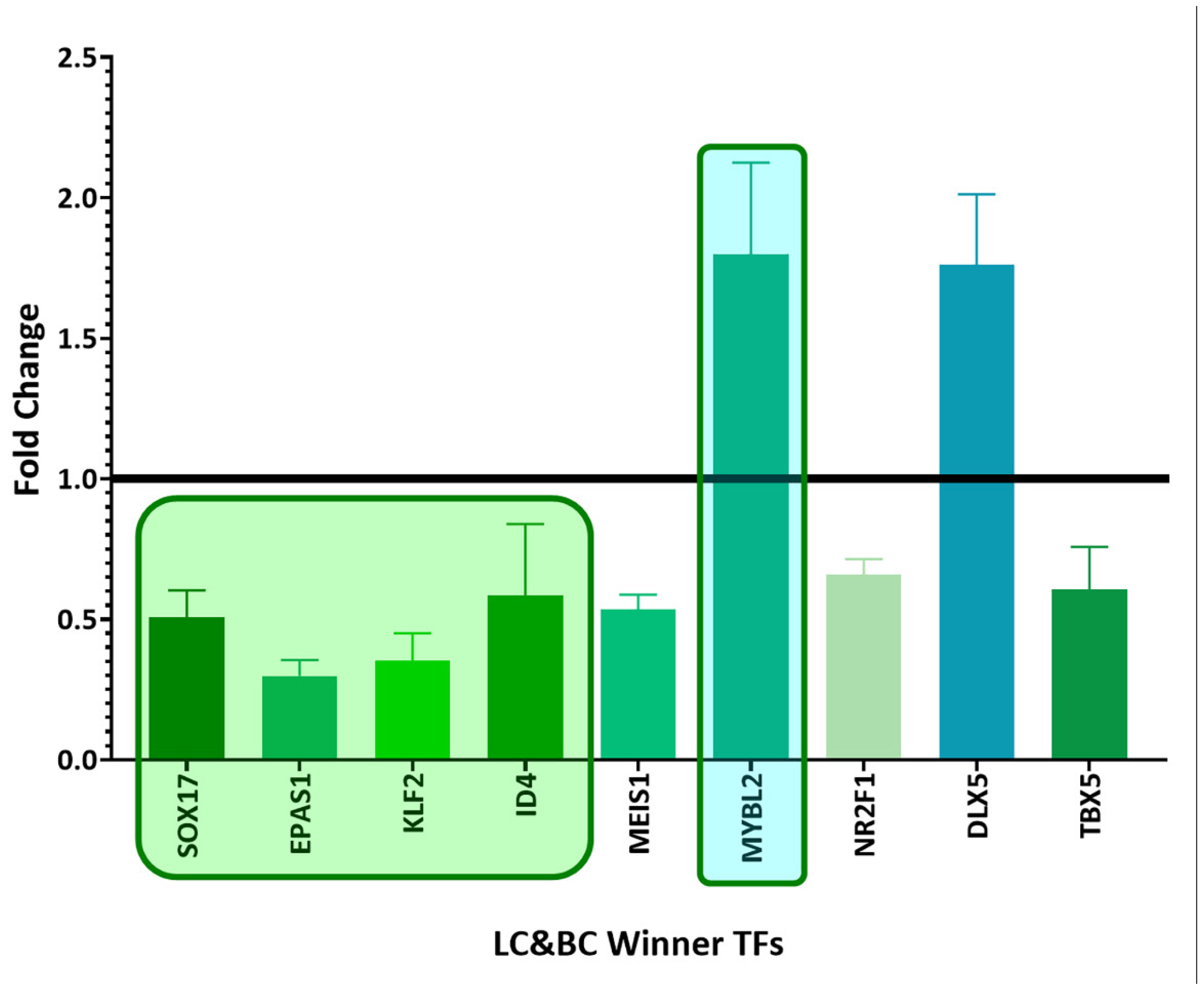
Biology, Free Full-Text
Recomendado para você
-
 Brain Test Nivel 367 Quiere tener grandes músculos16 junho 2024
Brain Test Nivel 367 Quiere tener grandes músculos16 junho 2024 -
 Brain Test Уровень 367 ответы (Он хочет большие мышцы)16 junho 2024
Brain Test Уровень 367 ответы (Он хочет большие мышцы)16 junho 2024 -
 Cabin Fever Word Search Puzzle – General Store of Minnetonka16 junho 2024
Cabin Fever Word Search Puzzle – General Store of Minnetonka16 junho 2024 -
 Ryuta Kawashima: The devil who cracked the dementia code, The Independent16 junho 2024
Ryuta Kawashima: The devil who cracked the dementia code, The Independent16 junho 2024 -
 Herpes Simplex Virus-1 in the Brain: The Dark Side of a Sneaky Infection: Trends in Microbiology16 junho 2024
Herpes Simplex Virus-1 in the Brain: The Dark Side of a Sneaky Infection: Trends in Microbiology16 junho 2024 -
 Prevagen Improves Memory - Regular Strength 10mg, 30 Capsules with Apoaequorin & Vitamin D & Prevagen 7-Day Pill Minder16 junho 2024
Prevagen Improves Memory - Regular Strength 10mg, 30 Capsules with Apoaequorin & Vitamin D & Prevagen 7-Day Pill Minder16 junho 2024 -
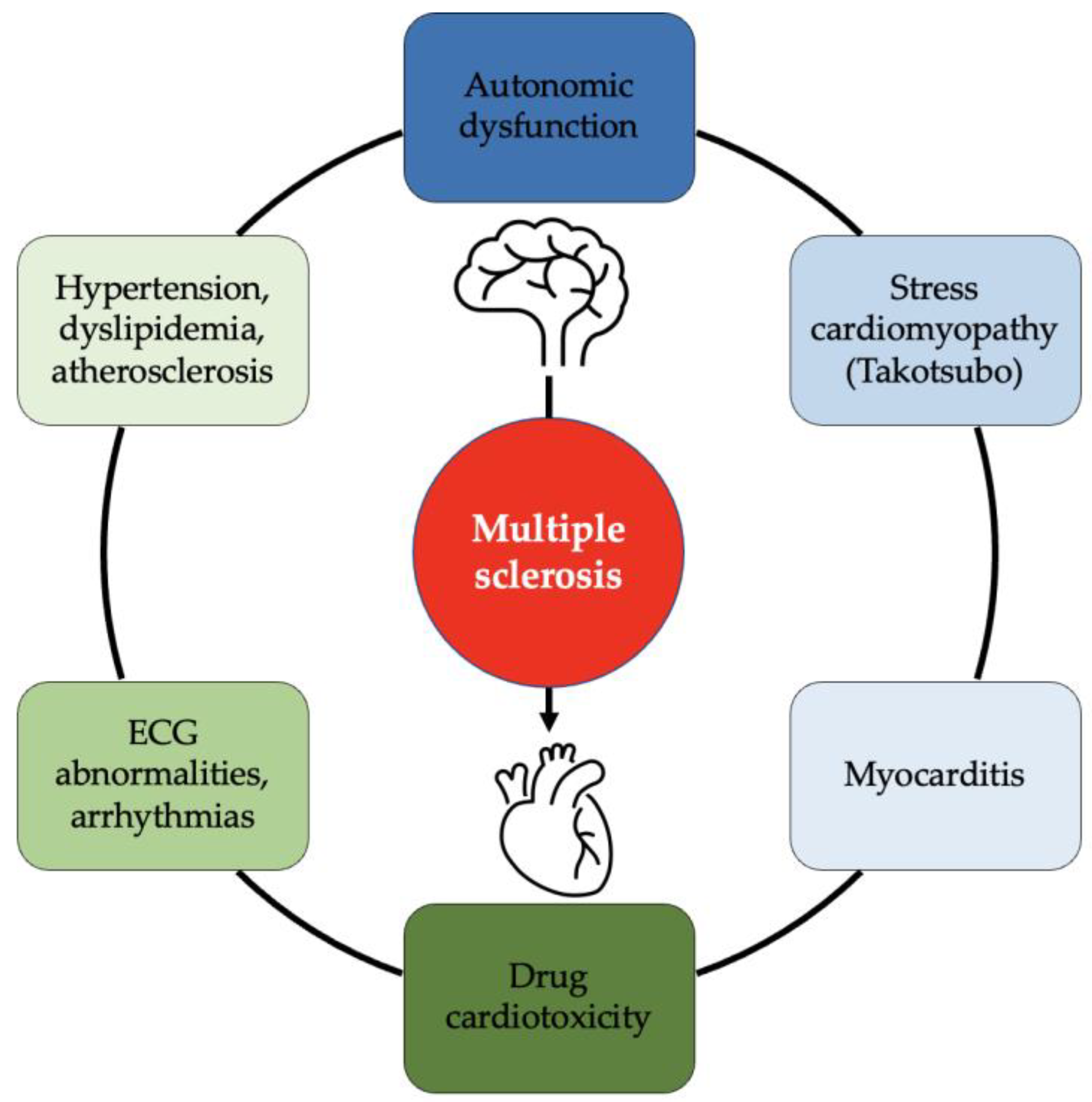 JCDD, Free Full-Text16 junho 2024
JCDD, Free Full-Text16 junho 2024 -
CBSE Class 12 Artificial Intelligence Question Paper 2023 with Answer Key (February 22, Set 4 - 367)16 junho 2024
-
 Stroop effect - Wikipedia16 junho 2024
Stroop effect - Wikipedia16 junho 2024 -
 Page 6 1,000+ Logo Alzheimers Care Pictures16 junho 2024
Page 6 1,000+ Logo Alzheimers Care Pictures16 junho 2024
você pode gostar
-
 House Adler Mashle Streetwear Hoodie16 junho 2024
House Adler Mashle Streetwear Hoodie16 junho 2024 -
 Pin em kits digitais gratuitos16 junho 2024
Pin em kits digitais gratuitos16 junho 2024 -
 Tiger Multiplier! na App Store16 junho 2024
Tiger Multiplier! na App Store16 junho 2024 -
 My Hero Academia Temporada 6 establece su fecha de estreno en16 junho 2024
My Hero Academia Temporada 6 establece su fecha de estreno en16 junho 2024 -
 Colorir os BISCOITOS DE NATAL em COQUINHOS16 junho 2024
Colorir os BISCOITOS DE NATAL em COQUINHOS16 junho 2024 -
Shiny Ultra Beasts - Roblox16 junho 2024
-
 Linda Hamilton Joins the Stranger Things Season 5 Cast - Netflix Tudum16 junho 2024
Linda Hamilton Joins the Stranger Things Season 5 Cast - Netflix Tudum16 junho 2024 -
 Desenhos para colorir, desenhar e pintar : Desenhos para colorir, moto16 junho 2024
Desenhos para colorir, desenhar e pintar : Desenhos para colorir, moto16 junho 2024 -
 Dragon Ball Super, Vol. 10 (10): 978197471526816 junho 2024
Dragon Ball Super, Vol. 10 (10): 978197471526816 junho 2024 -
 How to Play GameCube Games on the Wii • VGLeaks 3.0 • The best16 junho 2024
How to Play GameCube Games on the Wii • VGLeaks 3.0 • The best16 junho 2024